This article is more than 1 year old
Cycle fires up 50,000-core HPC cluster on Amazon EC2
Looking for drugs in all the right places
Cycle Computing is at it again, pushing the envelope on setting up HPC clusters that ride atop Amazon's EC2 compute clusters. This time, Cycle Computing has been tapped by protein simulation software-maker Schrödinger and drug-hunter Nimbus Discovery, which is in hot pursuit of drugs to cure Non-Hodgkin's lymphoma and obesity.
Schrödinger has expertise in creating molecular modeling software that can simulate proteins and their interactions with other chemicals floating in solution. In the absence of software modeling, the drug discovery process involves picking a protein in the body that is associated with a disease and seeing how it reacts with thousands, tens of thousands, hundreds of thousands, or millions of molecular snippets called ligands. This process is called screening in the wetware world, and it is obviously time consuming. It is also the kind of thing that, by its very nature, would lend itself to parallel processing if you could model proteins and ligands effectively and if you had a large amount of processing capacity to play with – at a decent price.
Simulating proteins and ligand snippets floating in the soup
The Glide software, as Ramy Farid, president at Schrödinger, explains it to El Reg, is a virtual screening package that can take a static protein molecule and simulate how different ligands would interact with that protein, which is called docking in the drug discovery lingo. The software has three different modes of operation, which allow researchers to play off time, compute resources, and the size of the data sets. Basically, you take different samples of some of the ligands in the simulation library and make some rough estimates about interactions in the coarsest mode, called High Throughput Virtual Screening, or HTVS.
Standard Precision, or SP mode, takes 10 times the resources to run, so you can generally only do it on a sample of the dataset, and the Extreme Precision mode takes 10 times more resources (or 100X the HVTS mode) to run. So generally, what companies do is take their best shot in HTCS mode, take 10 per cent of the ligands that might bind to a protein in an interesting way and run them through SP mode, and then take 10 per cent of these and run them through XP mode. When you are done, you have what you hope are compounds that might be suitable for development as a drug to affect proteins associated with a disease.
While this is how Schrödinger has worked with customers to help them try to find compounds that might make good drugs, this is not the proper way to do things, because if your sampling is too small at the beginning, you get false negatives and miss possible drugs.
"What we have been doing is cutting corners," explains Farid. "We didn't have a choice because we had to devise a protocol for screening that did less sampling than we wanted to do because we were limited by compute resources."
After being contracted by Nimbus Discovery to do Glide runs against proteins found to be interesting by the pharma startup, Schrödinger decided that what it needed to think about how it would use its own Glide software if it didn't have any compute capacity issues, and then get it running on the cloud. The company contracted Cycle Computing to build that cloud atop Amazon's EC2 compute cloud and configure the Glide images with its CycleCloud provisioning tool and manage the whole shebang with its CycleServer monitor for public clouds.
Now, rather than taking a subset of the 7 million interesting ligands in the Nimbus Discovery data set, Schrödinger ran the high-resolution SP docking routine against each and every one of those ligands, eliminating the possibility of any false negatives in that data set and also doing a much better level of simulation to boot. This revised docking protocol was not as simple as matching 7 million ligands against one rolled-up ball of protein, since the ligands themselves twist and bend in different configurations themselves. It was more like 21 million different ligand conformations that needed to be examined against the protein Nimbus Discovery was focused on.
The idea was to do all of this work in somewhere between two and three hours, when Schrödinger's internal cluster, which has 400 cores, would take about 275 hours to do the work using the old protocol – which did not give the same level of confidence as the new protocol would.
To that end, Cycle Computing fired up a cluster on Amazon that it nicknamed "Naga," which is Sanskrit for "cobra", among other things. The Naga virtual cluster was comprised of 6,742 Amazon EC2 instances with a total of 51,132 x86 cores and nearly 29TB of main memory. The server nodes were predominantly located in Amazon's US-East data center in Virginia, but spanned the globe thus:
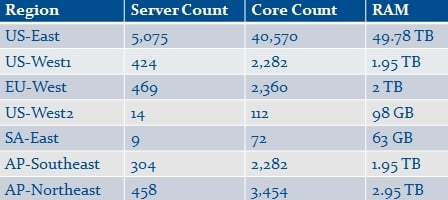
Feeds and speeds of the Naga cluster fired up by Cycle Computing
"HPC clusters are too small when you need them most, and too large the rest of the time," as Jason Stowe, Cycle Computing's CEO and founder, says wryly.
It is hard to say what a 50,000-plus core cluster would cost exactly, but depending on the configuration, you are looking at somewhere between $10m and $15m. But paying that kind of money is insane unless you have enough workloads to keep that cluster busy nearly all of the time.
Buying the capacity on Amazon through Cycle Computing makes a whole lot more sense. This Naga cluster, fully configured to run the job in three hours, cost $4,828 per hour, or about three orders of magnitude lower cost than having to buy a cluster to run the Nimbus Discovery job in the Glide software in-house.
Of course, that's not the end of it. Not only are ligands wiggly little fellows, but so are proteins, and in the Glide simulations, the proteins are held static because – you guessed it – there's not enough computing resources to let everything wiggle at once. And that, says Farid, is the next problem that Schrödinger wants to tackle. Allowing all of the molecules to twist around replicates how they really work inside of our bodies, and such simulations will result in better drugs being found more quickly. ®
